P-bodies are dynamic ribonucleoprotein granules found in the cell cytoplasm. They are primarily composed of translationally repressed mRNAs and proteins associated with mRNA decay machinery.
What are P-Bodies?
P-bodies (processing bodies) are evolutionarily conserved in eukaryotes and possess the characteristics of liquid droplets. An investigation on proteins related to mRNA decay pathways (decapping and breakdown) has found that P-bodies contain mRNA decay intermediates. This observation has led to an initial hypothesis that P-bodies are basically the cellular locations of mRNA decay.
It was later found that P-bodies are indispensable in regard to mRNA decay. In this context, a recent study has demonstrated that mRNA delay takes place in yeast strains lack P-bodies. Therefore, it is still uncertain what role P-bodies play in mRNA decay. One hypothesis is that they act as storage sites for translationally repressed mRNAs and inactive mRNA decay enzymes.
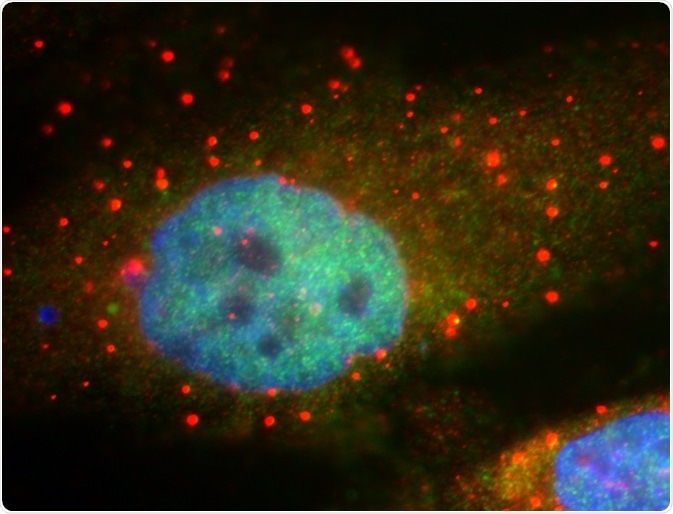
P-bodies are constitutively associated with proteins that are involved in translational repression and mRNA decay. These proteins include cytoplasmic deadenylase complex, decapping coactivator and enzyme, decapping activators, and 5’-3’ exoribonuclease.
In addition, RNA-binding proteins are also an integral part of P-bodies. According to a recent study, myosins are also associated with P-bodies, indicating a possible connection between P-bodies and cytoskeleton regulation.
Formation of P-bodies depends on liquid-liquid phase separation (LLPS), which means that these non-membrane cellular compartments are formed by the phase-separation from the cytoplasm. Recently, several fluorescence microscopy studies have revealed that a rapid exchange of components (proteins and RNA) occurs between P-bodies and the cytoplasm, suggesting that P-bodies have dynamic and liquid-like properties.
In addition, P-body dynamics that clearly indicate liquid-like properties are as follows: the spherical morphology of P-bodies due to surface tension as well as the fusion of two P-bodies to form a larger structure that maintains its original round shape. Hexanediol can reversibly dissolve P-bodies, which hinders weak intermolecular interactions.
Concentrations of many P-body-associated proteins with domains containing low complexity sequences and LLPS-associated RNA-binding activities play an essential role in the formation of new P-bodies as well as the maintenance of existing P-bodies. The presence of translationally repressed mRNA and polysome-associated RNA binding proteins is essential for maintaining P-bodies.
Regarding formation of new P-bodies, post-translational modifications of proteins, such as phosphorylation and ubiquitination, are known to have significant contributions. For instance, phosphorylation of decapping enzymes, Dcp1/2, is essential for P-body formation and localization. Moreover, ubiquitination of these enzymes plays an indispensable role in P-body assembly, either independently or by regulating phosphorylation of other P-body-associated proteins. Besides these physiological modifications, certain stress conditions, such as glucose starvation and osmotic stress, are also known to increase the formation of new P-bodies.
Functions of P-Bodies
P-bodies play an essential role in regulating mRNA metabolism. Initially it was thought that P-bodies participate in mRNA decay; however, recent studies using mRNA single-molecule imaging system have clearly demonstrated that mRNA decay events occur in yeast cytoplasm rather than in P-bodies.
In addition, these studies have also revealed that mRNA decay products are present throughout the cytoplasm rather than being stored in P-bodies. All these findings clearly indicate that P-bodies are not directly involved in mRNA decay; they may act as reservoirs to store translationally repressed mRNAs and inactive decapping enzymes.
Stress-induced activation of P-bodies can lead to the inhibition of translation initiation. P-bodies are also known to store proteins and mRNAs associated with nonsense-mediated decay, a special type of mRNA quality control process for degrading mRNAs that terminate translation aberrantly. Moreover, in some cases, P-bodies also contain proteins and microRNAs related to the microRNA repression pathway.
According to many experimental confirmations, mRNAs recirculate between P-bodies and polysomes. One possible mechanism is that mRNAs that cannot participate in translation interact with decapping machinery, and the formed complex is transported to P-bodies from polysomes. Such aggregation of translationally repressed mRNAs inside P-bodies may be essential for maintaining proper amounts of mRNAs within polysomes as well as regulating effective translation of mRNAs into proteins.
Sources
- https://www.uniprot.org/locations/SL-0230
- www.sciencedirect.com/…/p-bodies
- https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3428773/#A012286C41
- https://pubs.acs.org/doi/abs/10.1021/acs.biochem.7b01162
Further Reading
- All RNA Content
- What is RNA?
- RNA Structure
- Types of RNA: mRNA, rRNA and tRNA
- RNA Synthesis
Last Updated: Sep 12, 2018

Written by
Dr. Sanchari Sinha Dutta
Dr. Sanchari Sinha Dutta is a science communicator who believes in spreading the power of science in every corner of the world. She has a Bachelor of Science (B.Sc.) degree and a Master's of Science (M.Sc.) in biology and human physiology. Following her Master's degree, Sanchari went on to study a Ph.D. in human physiology. She has authored more than 10 original research articles, all of which have been published in world renowned international journals.
Source: Read Full Article


