The electron transport chain is comprised of a series of enzymatic reactions within the inner membrane of the mitochondria, which are cell organelles that release and store energy for all physiological needs.
As electrons are passed through the chain by a series of oxidation-reduction reactions, energy is released, creating a gradient of hydrogen ions, or protons, across the membrane. The proton gradient provides energy to make ATP, which is used in oxidative phosphorylation.
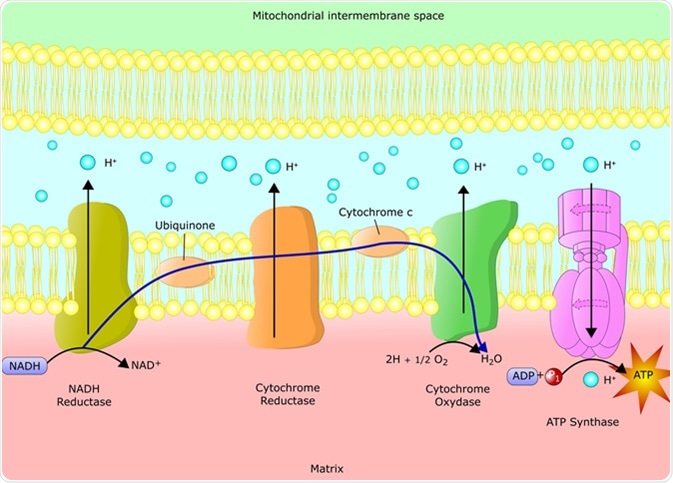
The reactions of the electron transport chain are carried out by a series of membrane proteins and organic molecules. They are arranged in four complexes. In eukaryotes, the electron transport chain is located in the inner mitochondrial membrane. In prokaryotes, it is located within the plasma membrane.
Electrons move through the electron transport chain from a higher to lower energy state. Energy release moves protons through channels in the membrane proteins, moving them into the inner membrane space. This leads to a buildup of positively charged protons, which creates an electrical potential across the membrane.
The reactions of the electron transport chain involve several large membrane protein complexes within the inner mitochondrial membrane. Some are described below.
The NADH Dehydrogenase Complex
The NADH dehydrogenase complex (Complex I) contains more than 40 polypeptides. It transfers electrons from NADH to coenzyme Q10. The reaction begins when NADH binds to Complex I, transferring two electrons to the flavin mononucleotide (FMN) prosthetic group, resulting in the formation of FMNH2. The electrons are then transferred through iron-sulfur clusters to coenzyme Q10. The change in redox state of the protein induces a conformational change, causing the four hydrogen ions to be pumped into the inner membrane space. Four protons are thus transported across the membrane in the reaction.
Succinate Dehydrogenase (Complex II)
Succinate dehydrogenase, also known as succinate-CoQ reductase, receives electrons into the quinone pool from succinate and transfers them to to Q. Complex II has four subunits. Complex II operates parallel to Complex I. However, no protons are transported into the intermembrane space. This enzyme also takes part in the tricarboxylic acid (citric acid) cycle as well.
The Cytochrome b-c1 Complex
The cytochrome b-c1 complex (Complex III), has 11 polypeptide chains and functions as a dimer, and is also known as coenzyme Q: cytochrome c-oxidoreductase or cytochrome c reductase. Three heme groups are found within each monomer, bound to cytochromes and an iron-sulfur protein. The function of the b-c1 complex is via a Q-cycle mechanism. It catalyzes the reduction of cytochrome c by the oxidation of coenzyme Q while pumping 4 protons from the mitochondrial matrix to the intermembrane space. Mutations of Complex III are associated with exercise intolerance and some multisystem disorders.
Cytochrome c Oxidase
Cytochrome c oxidase is the last step in the electron transport chain. It functions as s dimer, with each monomer containing 13 different polypeptide chains, including two cytochromes and two copper atoms. It accepts two electrons from two cytochrome c molecules and passes them four at a time to oxygen. Mutations of cytochrome c oxidase can lead to severe metabolic disorders. The cytochrome oxidase reaction uses about 90 percent of the oxygen taken up by most cells.
Uncoupling
Electron transport can be uncoupled from ATP synthesis through the use of certain agents or some natural processes. Some specialized fat cells, known as brown fat, uncouple the electron transport chain in order to dissipate the energy as heat. This is accomplished through a transport protein that moves protons down the electrochemical gradient, bypassing ATP synthase. The cells oxidize their fat stores rapidly, producing heat. Hibernating animals and newborn human babies have brown fat.
Sources
- Molecular Biology of the Cell, 4th Edition, https://www.ncbi.nlm.nih.gov/books/NBK26904/
- The respiratory-chain NADH dehydrogenase (complex I) of mitochondria, onlinelibrary.wiley.com/doi/pdf/10.1111/j.1432-1033.1991.tb15945.x
- The cytochrome bc-1 complex: function in the context of structure, www.annualreviews.org/…/annurev.physiol.66.032102.150251
- http://www.life.illinois.edu/crofts/bioph354/succ_dh.html
Further Reading
- All Mitochondria Content
- Analyzing Mitohormesis
- Mitochondrial DNA Organization and Replication Steps
- Mammalian Mitochondrial DNA (mtDNA) Transcription
- Mitochondrial DNA (mtDNA) Translation in Eukaryotes
Last Updated: Feb 26, 2019

Written by
Dr. Catherine Shaffer
Catherine Shaffer is a freelance science and health writer from Michigan. She has written for a wide variety of trade and consumer publications on life sciences topics, particularly in the area of drug discovery and development. She holds a Ph.D. in Biological Chemistry and began her career as a laboratory researcher before transitioning to science writing. She also writes and publishes fiction, and in her free time enjoys yoga, biking, and taking care of her pets.
Source: Read Full Article


